Visualisation of Blood Group Antigens
At the ITM, a database for all proteins that exist on the surface of erythrocytes is being developed. The database will contain all proteins that carry blood group antigens, including all known variants. This new resource combines available information about genetic polymorphisms with information on the function and 3D structure of the proteins. Unknown structures are predicted with the deep-learning algorithm AlphaFold. Computational molecular dynamics simulations allow us to predict the effect of blood group variations on the structure of the proteins, and consequently their effect on the erythrocyte surface.
An example for the benefit of combining genetic and structural information is the blood group protein Kell. The different antigens on the surface of this protein can cause massive adverse immune reactions in pregnant women or after blood transfusions. Additionally, the Kell protein can occur both alone and in a covalent complex with another blood group protein called XK. The predicted structures of Kell alone vs. Kell bound to XK illustrate a substantial difference in the position of the Kell protein relative to the erythrocyte surface (Fig. 1).
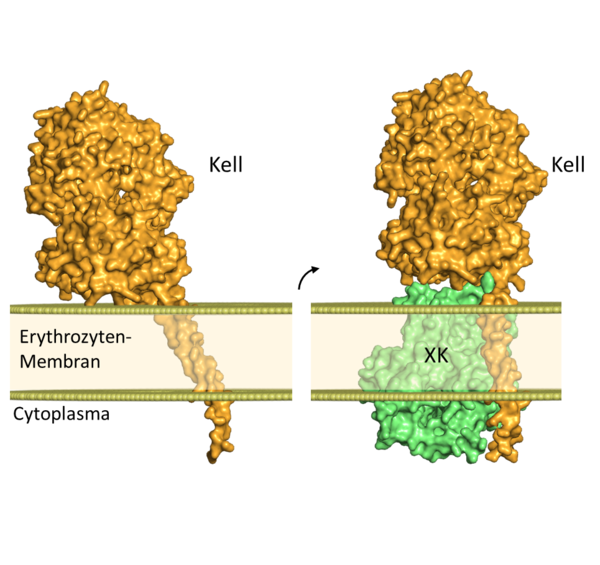
Figure 1: Predicted structure and orientation of the blood group protein Kell int the erythrocyte membrane; Left: Kell alone, right: Kell in complex with XK. Binding to the XK protein causes a change in the position of the Kell protein in the membrane. The intracellular termini of the proteins are hidden in this figure due to the low confidence scores of the AlphaFold prediction in this region.

